by Caleb Sooknanan ’20
Influenza is a common viral infection that attacks the respiratory system. Major outbreaks occur due to antigenic changes in the influenza A virus, which is when virus strains from separate hosts combine to form different strains with a mixture of surface antigens. Unfortunately, the mechanism behind this replacement, or antigenic shift, remains misunderstood. Dr. Yuki Furuse and his team of researchers at the Tohoku University Graduate School of Medicine used a theoretical susceptible–exposed–infectious–recovered (SEIR) model to simulate and understand the replacement mechanism. This was also done to analyze factors in the virus and host populations that could affect antigenic shift.
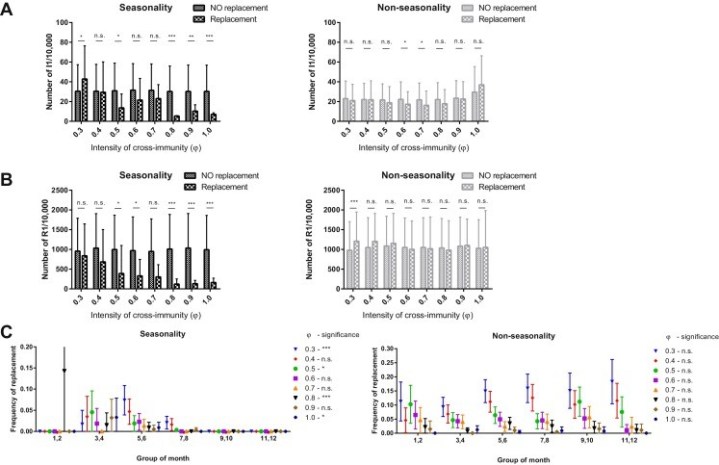
The model was stochastically created based on a previously circulating viral strain and a new strain. The results suggested that antigenic replacement was more likely to occur in tropical regions than in temperate regions due to the virus’s higher reproductive number. Based on the higher concentration of viral antigens present in tropical regions, the researchers theorized that replacement was not affected by the timing of the new strain’s appearance. The data chart (Figure 1) shows the “number of I1,” the current epidemic magnitude from the existing strain, and the “number of R1,” the herd immunity against the existing strain. I1 and R1 were lower for the emerging strain, thereby suggesting that replacement occurs only when an emerged strain appears at an epidemic’s initial stage.
Even with these results, the researchers suggested that it could be difficult to detect new viral strains in temperate regions with low circulation levels. A very small percentage of circulating strains can be detected and therefore categorized. However, emerging strains in tropical regions or strains imported from a temperate region to a tropical region are more likely to be detected because of high replacement probabilities.
Limitations to the study included its lack of age-specific parameters. Also, the stochastic model set different infection rates and immunity for each person, thus affecting the overall results. Further investigations are needed for scientists to understand viral transmission dynamics and develop potential strategies to diminish the effects of seasonal and pandemic influenza.
References:
- Y. Furuse, et al., Mechanisms of replacement of circulating viruses by seasonal and pandemic influenza A viruses. International Journal of Infectious Diseases 51, 6-14 (2016). doi: 10.1016/j.ijid.2016.08.012
- Image retrieved from: http://www.sciencedirect.com/science?_ob=MiamiCaptionURL&_method=retrieve&_eid=1-s2.0-S1201971216311390&_image=1-s2.0-S1201971216311390-gr4.jpg&_cid=272991&_explode=defaultEXP_LIST&_idxType=defaultREF_WORK_INDEX_TYPE&_alpha=defaultALPHA&_ba=&_rdoc=1&_fmt=FULL&_issn=12019712&_pii=S1201971216311390&md5=370c120c4cd0b4114d2fe52cde60ec21


